Research
The Tang Lab develops computational models to understand biological systems across scales and modalities. We focus on building predictive, interpretable, and autonomous AI models that analyze and forecast cellular, tissue, and organismal behavior, enabling reliable in silico experiments that complement wet-lab efforts. Our overarching goal is to extract mechanistic insights from data—spanning cell biology, multi-modality biology, neuroscience, cancer, development and aging, and brain–computer interfaces.
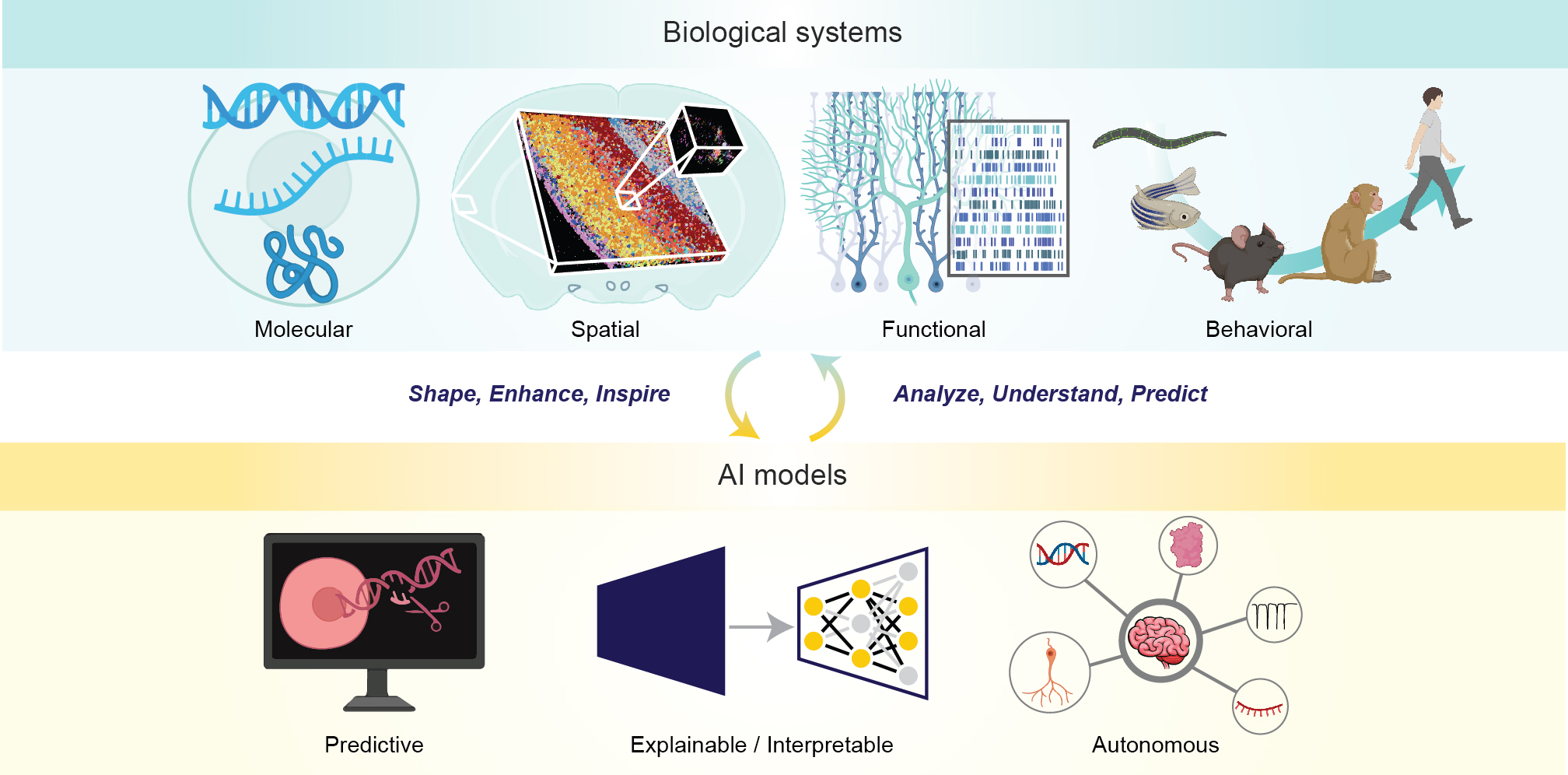
We aim to bridge large-scale and limited-scale cellular measurements with the underlying mechanisms that govern biological function. Our research centers on generic and powerful AI methods for biology and medicine. Briefly, our major research questions include:
Decoding Cancer with AI
At the molecular level, how do genomic and epigenomic alterations reorganize chromatin architecture and rewire gene-regulatory networks? Moreover, how are these intracellular perturbations conveyed through intercellular communication to promote the spatiotemporal progression of tumor growth?

Decoding Brain with AI
At the functional level, how can we establish correlations between specific genes, specific cells—especially neurons and neuronal ensembles that exhibit unique electrical dynamics—and diverse animal behaviors and brain disease?

Decoding Aging with AI
Across the lifespan and at system level, how do genetic, epigenetic, proteostatic, and metabolic trajectories—shaped by niche, immune, and environmental cues—drive cellular and tissue aging? Which circuits are causal, reversible, and targetable, and how can AI disentangle age, disease, and environment to predict resilience and response to interventions, and prioritize rejuvenation interventions?

Building Virtual Cells
On a silicon chip, how can we develop autonomous, data-driven AI approaches to bridge the gap between large-scale/small-scale cellular measurements and the contextualized mechanism of cellular and system behavior? And how can we design approaches to help the broader biologists in its easiest way?
Building Self-driving Lab
At the bench, how can we create a self-driving experimental platform that connects robots, instruments, and AI to design, execute, and learn from experiments in a closed loop? How do we translate hypotheses from our virtual cells into executable protocols, rapidly iterate with active learning, and make the entire system accessible and safe for biologists through robust interfaces, reproducibility, and traceability?
Funding Support
We acknowledge the generous funding support from UBC Michael Smith Laboratories and Department of Computer Science, UBC AI and Health Network, The Digital Research Alliance of Canada, Canada Research Chairs, NSERC Discovery, Canada Foundation for Innovation, and BC Knowledge Development Fund.